Motivated Proteins
A Web Facility for Studying Small Hydrogen-Bonded Motifs
What is Structure Motivator?
Structure Motivator is a desktop application for studying the range of structures represented by small three-dimensional protein motifs. It gives off-line access to the same motifs as in the Motivated Proteins web application and allows the user to refine them further. In 2021 it was updated to version 2, incorporating the same approx. 4000 proteins available in Motivated Proteins 2.
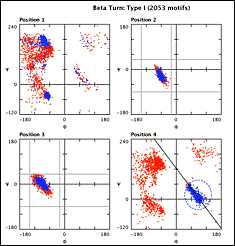
Structure Motivator presents a group of motifs as points in a plot of dihedral angles. In addition to φψ plots, there are also options to view side-chain χ1 angles and the dihedral angles bounding each peptide bond. The key feature of the application is the ability to make selections of motifs from these plots, interactively with the mouse cursor and/or on the basis of the presence or absence of specific residues within the motif. Details of selected residues may be displayed in a list, from which they may be visualized in the Jmol 3D protein-viewer. Selections may be exported to spreadsheet or reimported for further analysis, and print and graphics functions are available.
A description of Structure Motivator has been published in BMC Structural Biology (2012) 12:26.
Download
Structure Motivator 2 version 1.1 was released on 12th May 2022. It requires Java to be installed on your computer (not your web browser) in order to run. It is not available for mobile devices.
- Mac OS X (Mac dmg — in case of problems try the Unix double-clickable jar file.)
- Windows 32-bit (zipped exe)
- Windows 64-bit (zipped exe — in case of problems try the Unix double-clickable jar file.)
- Unix/Linux (zipped jar — also usable on Windows and Mac)
- Structure Motivator Manual (pdf — also included in the application distributions)
- Additional Motif and Demo Files (zipped folder)
PreMotivator 2 is a small utility for preparing input files for the Structure Motivator 2 application. The user can specify the main-chain dihedral angles (φ and ψ) for each residue of a short peptide (up to 9 residues in length) and the application will search for matching instances in an embedded database of approx. 4000 protein chains. The results may be exported as files suitable for loading into Structure Motivator 2. It is included in the Structure Motivator distributions.
