This glossary presents the definitions used for the small weakly-bonded motifs in the database underlying the Motivated Proteins web application. Individual motif categories are primarily defined by the pattern of hydrogen bonding or, in some cases, ionic bonding or potential for ionic bonding. The subcategories may be partially defined by alternative motif sizes or residue identities, implicit in the category definitions, but in many cases the key subdivisions relate to the dihedral angles found at specific positions in the motifs. This is explained in more detail below.
Hydrogen Bonding
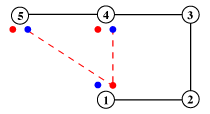
The hydrogen-bonding patterns of motifs are represented by diagrams of the type shown in the example below. Alpha-carbons are represented as numbered open circles, and and the associated main-chain NH and CO of residues involved in hydrogen bonding are represented as smaller filled circles, coloured blue and red, respectively. The corresponding side-chain hydrogen bonding atoms are represented as un-numbered squares, coloured according to the cpk scheme, as before (sulphur is yellow). Hydrogen bonds are shown as red broken lines, ionic bonds as broken black lines.
Dihedral Angles
The dihedral angles φ and ψ — the angles of rotation about the bonds to the α-carbon atom — describe the main chain conformation. χ1, χ2 etc. are the successive side chain rotation angles (Asp is illustrated).

Because of steric repulsion, only certain combinations of φ and ψ are allowed, as indicated by the Ramachandran plot, below. These are the well-known αR and βR conformations, together with the less-abundant corresponding left-hand forms.
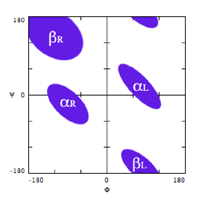
The χ1 angle of the side-chains involved in motif hydrogen-bonding — those of Asp, Asn, Ser, Thr and Cys — are also constrained, and have values of either approximately 60°, 180° or –60°.
The default ranges used to define the φ and ψ angles for the motifs are:
- αR: φ = –140° to –20°; ψ = –90° to +40°
- αL: φ = +20° to +140°; ψ = –40° to +90°
- βR: φ = –140° to –20°; ψ = +80° to +180°
- βL: φ = +20° to +140°; ψ = –180° to –80°
The χ1 angles quoted allow for 40° above and below the stated value.
Motif enantiomers
Where a motif is defined by hydrogen-bonds involving only certain positions, alternative combinations of dihedral angles may be possible at other positions. In those cases where the motif contains a loop of two unbonded residues (c.f. positions 2 and 3 in the diagram above) two types of enantiomers are found, leading to four possible structures. First there is the αR-αR structure (generally the most common and also referred to as type I) and the corresponding αL-αL mirror image (type I'):
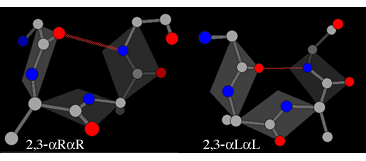
In each of these the plane of the peptide can flip through 180° while still maintaining the hydrogen bond. The flipped form of the αR-αR structure is βR-αL (also known as type II), whereas the flipped form of the αL-αL structure is βL-αR (also known as type II').
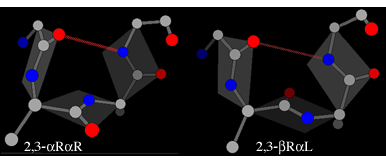
The illustrations also introduce the shorthand used to indicate which are the enantiomeric residues in a motif.
Alternative nomenclature
The numbering scheme used above is based on the standard one used for β-turns — components of many of the other motifs presented here. However for compound motifs that include both a β-turn and a β-bulge (the β-bulge loop and the β-turn) a conflict arises between different numbering schemes used for the two components. In these cases both are presented.